mRNA vaccines and drugs
The mRNA vaccine broke the traditional immune activation mode of inactivated and attenuated vaccines, and innovatively used human cells to produce antigens to activate specific immunity, which was brilliant during the COVID-19 epidemic. The application prospects of mRNA vaccines and drugs are very broad. In addition to being used for preventing infectious diseases, treating tumors and immune diseases, they can also be applied in various biopharmaceutical platform technologies, such as protein supplementation therapy, cell therapy, antibody drugs, gene editing, and so on. In the post COVID-19 era, mRNA vaccines still have the potential for substantial growth.
Preparation of mRNA immune antigen antibody
mRNA has the potential to encode almost all proteins. mRNA vaccine technology can be applied to antigen preparation to solve the difficulties in protein preparation, such as long preparation cycles, complex production processes, and poor flexibility in customized production of traditional protein antigens. mRNA, as a platform technology, has the advantages of simple production process, short production cycle, and high flexibility. It can provide customers with antigen preparation faster and more cost effectively, and conduct rapid target feasibility verification.
Hzymes Biotech provides one-stop mRNA vaccine&drug development concept validation services to help customers quickly verify the feasibility of drug targets
- Experienced team has successfully delivered multiple concept validation projects
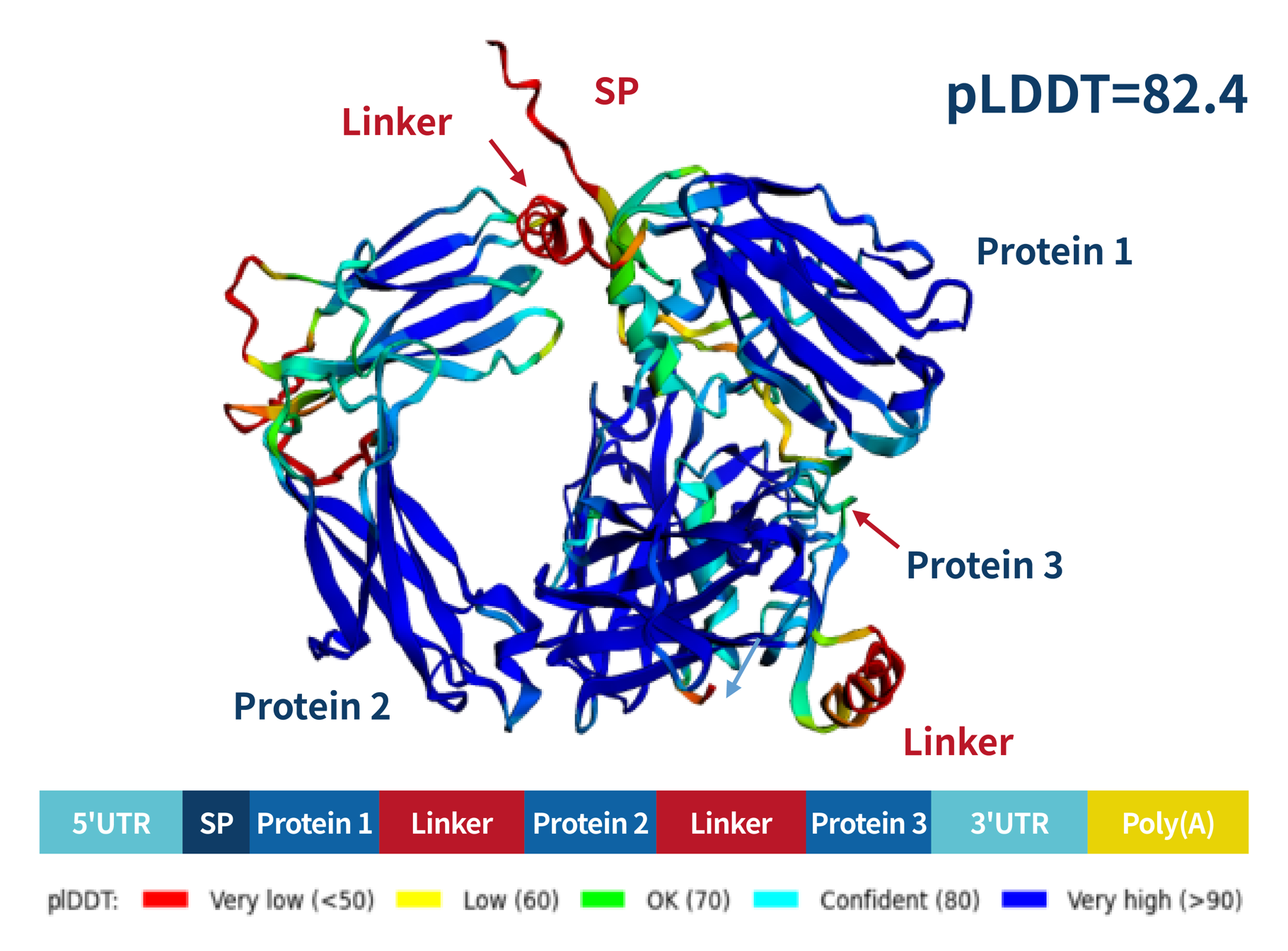
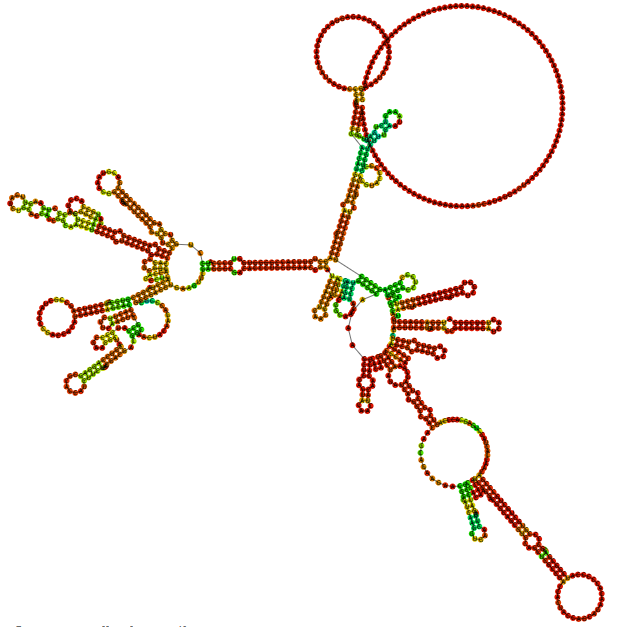
- Diversified tools are used to predict the secondary structure of proteins using AlphaFold, providing guidance for customers' antigen design and mRNA sequence design.
- Our independently established mRNA technology and application platform, comprehensive quality system, and full process services from sequence design, mRNA LNP preparation to in vivo biological evaluation.








Service category | Duration (working days) | Deliver |
Sequence design | 5 | 10 candidate sequences |
Plasmid construction and amplification | 30 | Plasmids and sequence files, providing synthesized raw plasmids (1-2 μ g) |
mRNA drug substance synthesis | 5 | mRNA Drug Substance Detection Report |
mRNA cell translation level detection | 10 | Translation Proficiency Test Report |
LNP encapsulation | 10 | 1mg LNP |
In vivo translation level detection (optional) | 20 | Test report |
In vivo cellular immune testing (optional) | 10 | Test report |
Humoral immune testing (optional) | 20 | Test report |
In vivo translation level+humoral immune testing (optional) | 20 | Test report |
In vivo translation level+humoral immune detection+cellular immune detection (optional) | 20 | Test report |


T7 promoter+transcription start sequence | IVT production | |
saRNA-EGFP(AU,UTP) | TAATACGACTCACTATAATG GGCGGCGCAT | Normal,1:181 |
TAATACGACTCACTATAATG CGGCGCAT | No output | |
TAATACGACTCACTATAAT CGGCGCAT | No output |


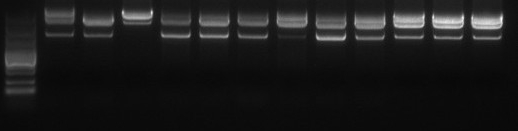
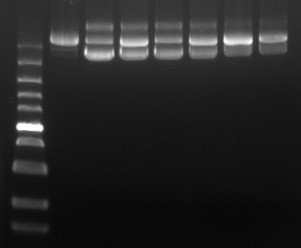
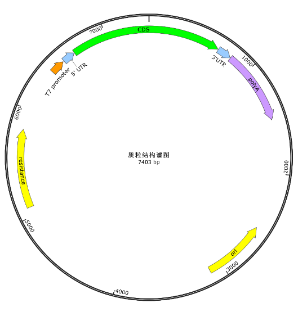
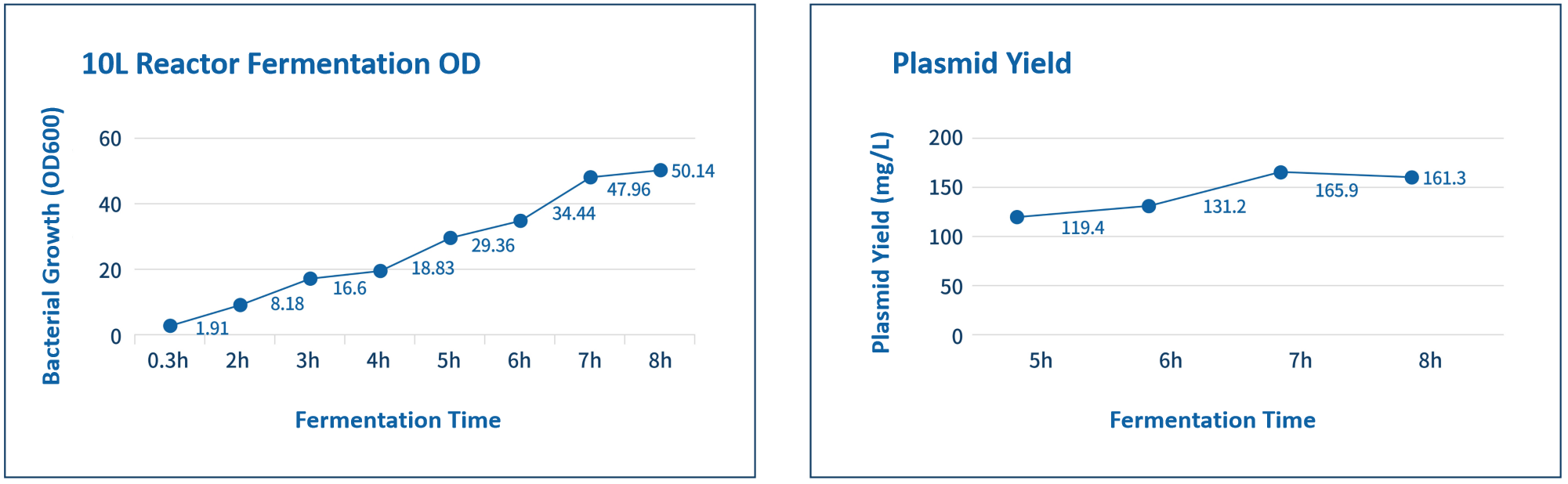
Quality control items | Number | Testing items | Test method |
Bacterial seed bank inspection | 1 | Bacterial morphology | Microscopic observation |
2 | Plasmid restriction enzyme digestion map | Restriction endonuclease digestion | |
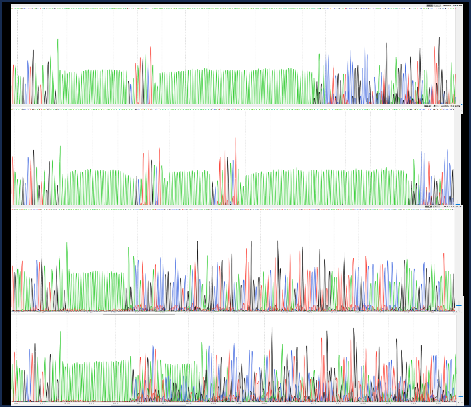
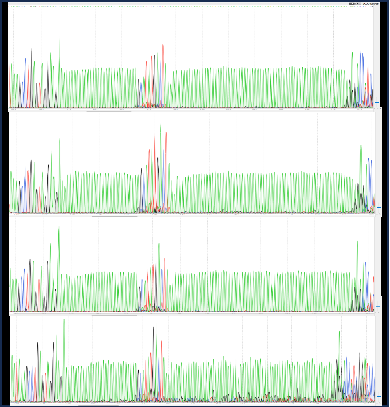
3 | Target gene sequencing | Sanger Sequence | |
4 | Sequencing of other components | Sanger Sequence | |
5 | Bacterial identification | Cultivation method/bacterial staining/biochemical reaction | |
6 | Antibiotics resistance | Cultivation method | |
The stability of strain passage | 1 | Plasmid sequence size | Electrophoresis/Sanger Sequencing |
2 | Accuracy of plasmid sequence | Sanger Sequence | |
3 | Plasmid restriction enzyme digestion map | Restriction endonuclease digestion | |
4 | Plasmid copy number | ddPCR | |
5 | Passage restrictions | Cultivation method | |
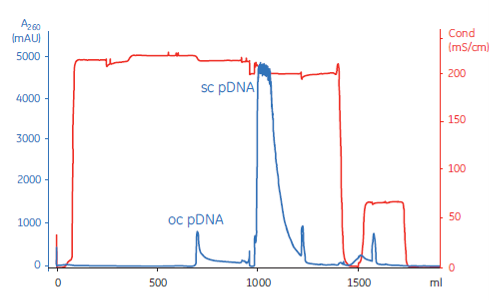
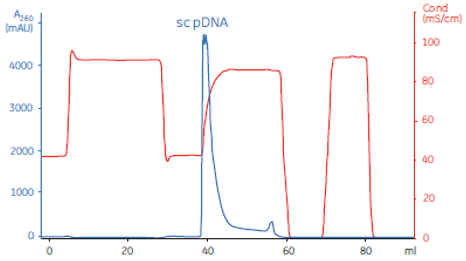
Quality control of supercoiled plasmids | 1 | Appearance | Visual inspection method |
2 | PH value | pH meter | |
3 | Purity (A260/A280) | Ultraviolet spectrophotometer | |
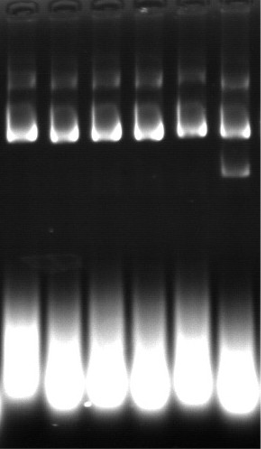
4 | Identification of restriction endonucleases | Agarose gel | |
5 | Sequencing identification | Sanger Sequence | |
6 | Concentration | Spectrophotometry | |
7 | DNA homogeneity | Electrophoresis or HPLC | |
8 | Host protein residues | Elisa | |
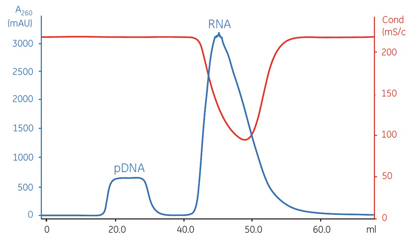
9 | RNA residue | Electrophoretic method | |
10 | Host DNA residue | QPCR | |
11 | Microbial limit/sterility | Membrane Filtration | |
12 | Endotoxin | TAL |
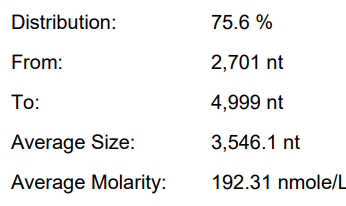




WT-T7 (Customer System)
Yield: 191.9 μ g/20 μ L reaction
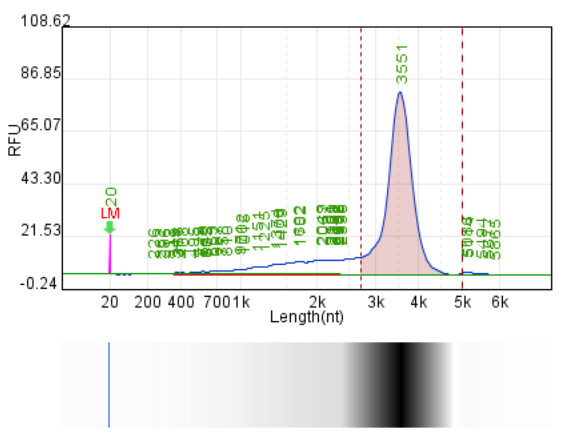
Integrity: 75.6%
Replace M6/optimize enzyme dosage
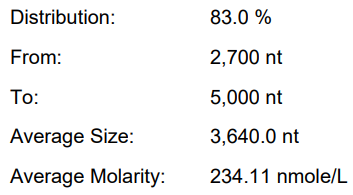
Yield: 176.8 μ g/20 μ L reaction
Integrity: 83.0%
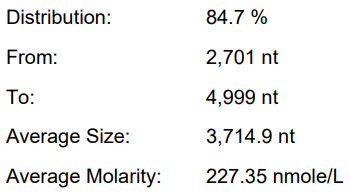
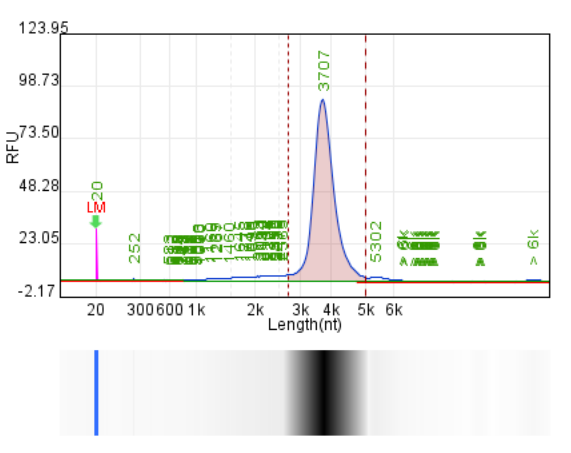
Database analysis and buffer optimization
Yield: 240.1 μ g/20 μ L reaction
Integrity: 84.7%
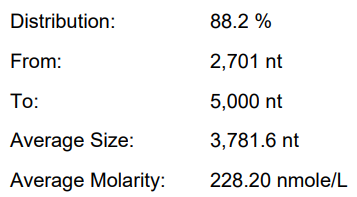
Use enhancers (final IVT system)
Yield: 203.8 μ g/20 μ L reaction
Integrity: 88.2%
Product release testing
Quality inspection items | Quality inspection method | Results Display |
Concentration | Nanodrop method | 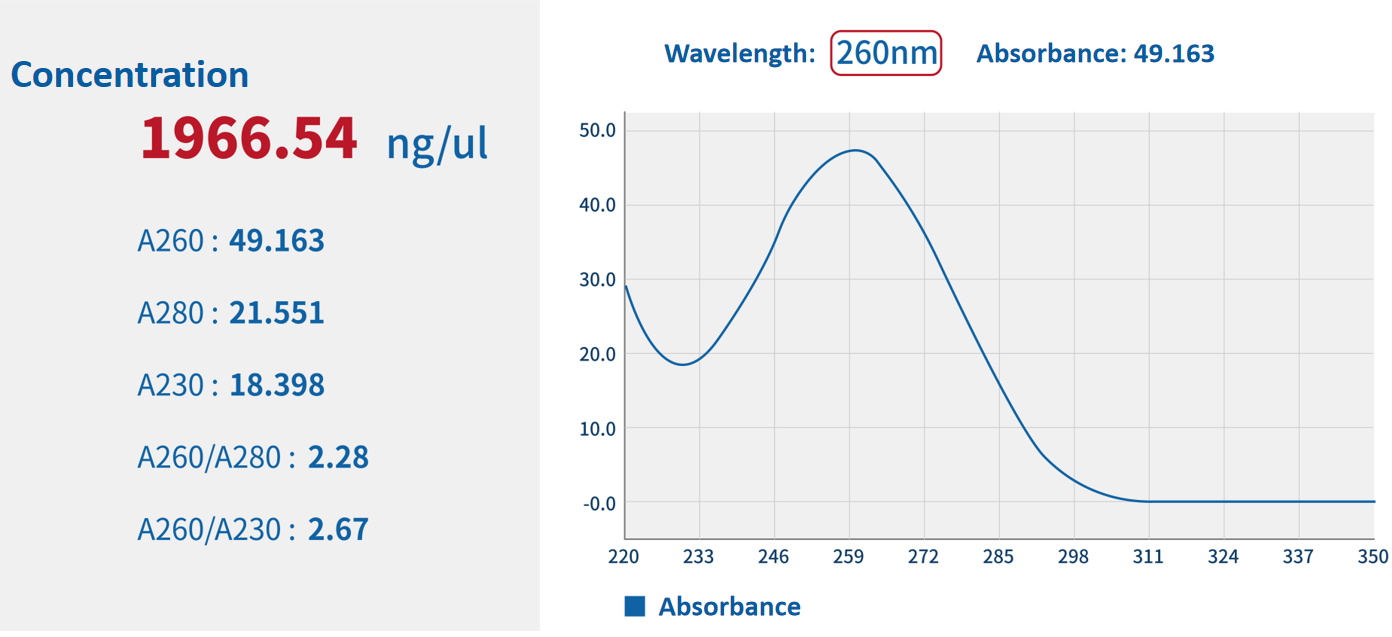 |
Ribogreen | 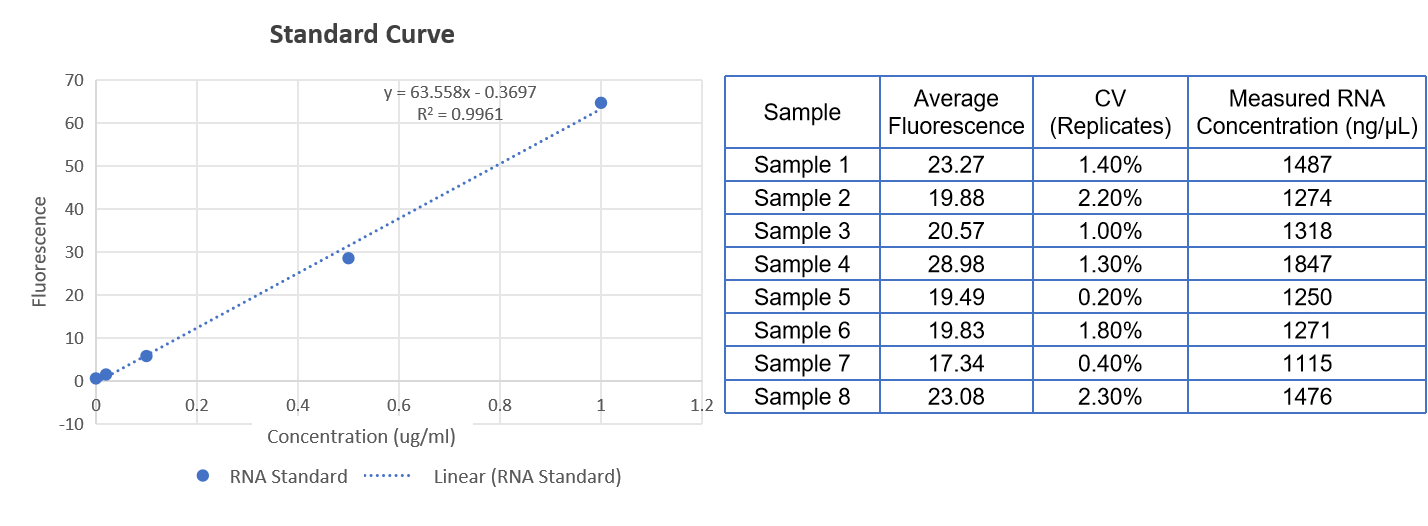 | |
Integrity | CE | 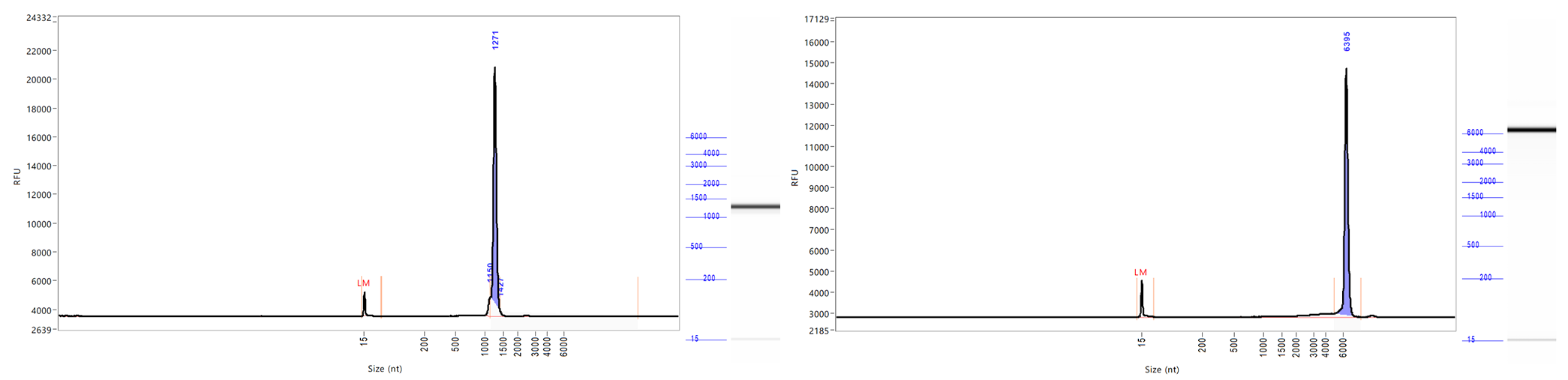 |
SEC-HPLC | 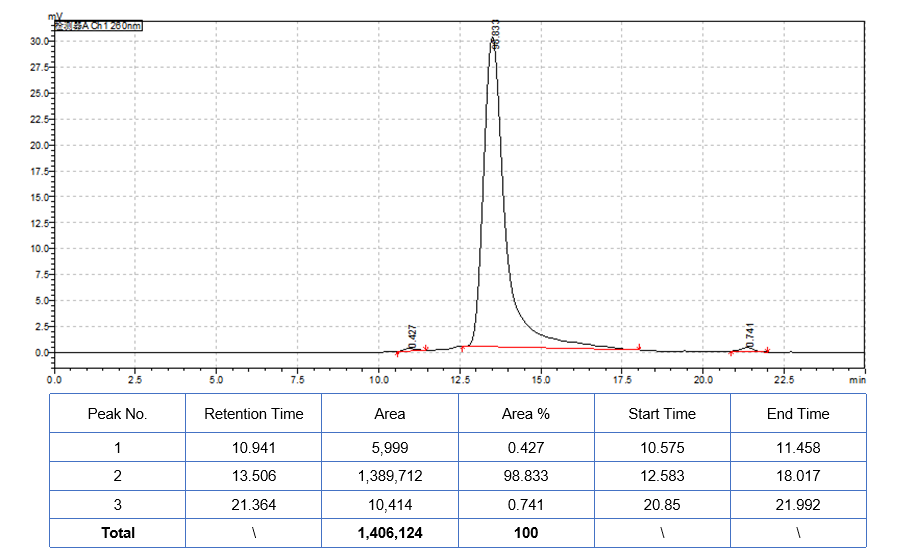 | |
DNA template residue | qPCR | 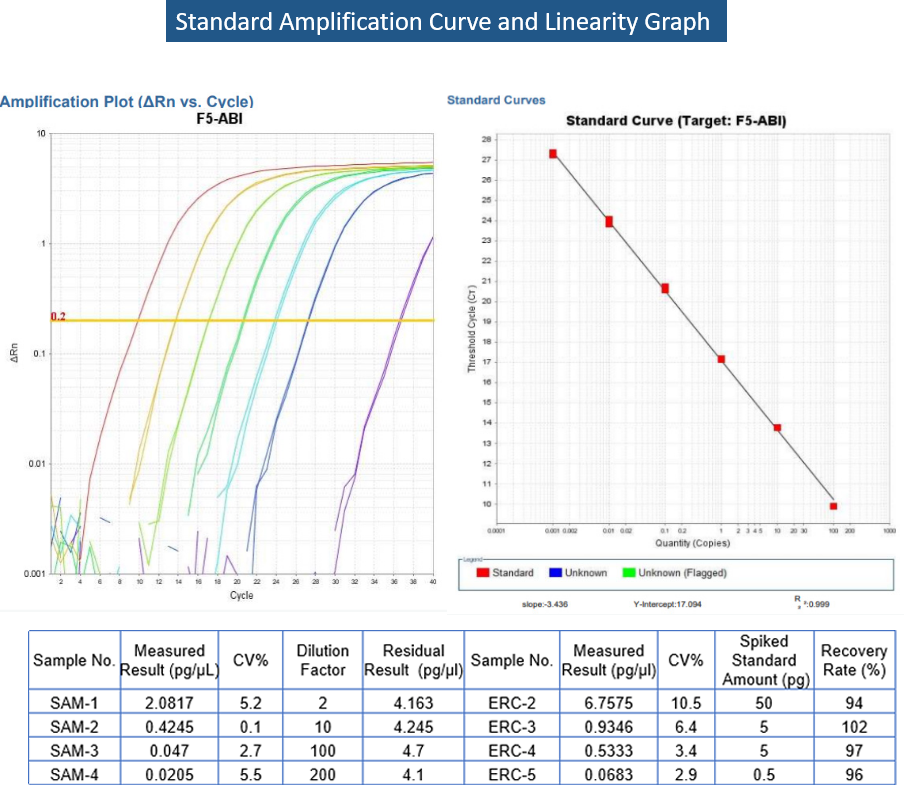 |
Capping efficiency | LC-MS | 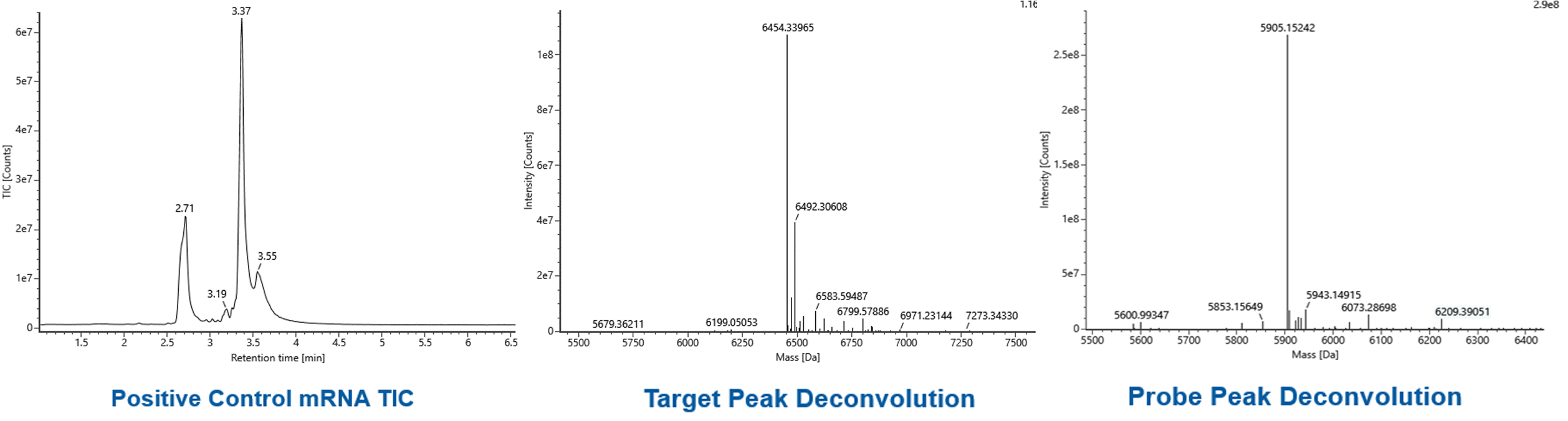 |
Poly (A) tail length | LC-MS | 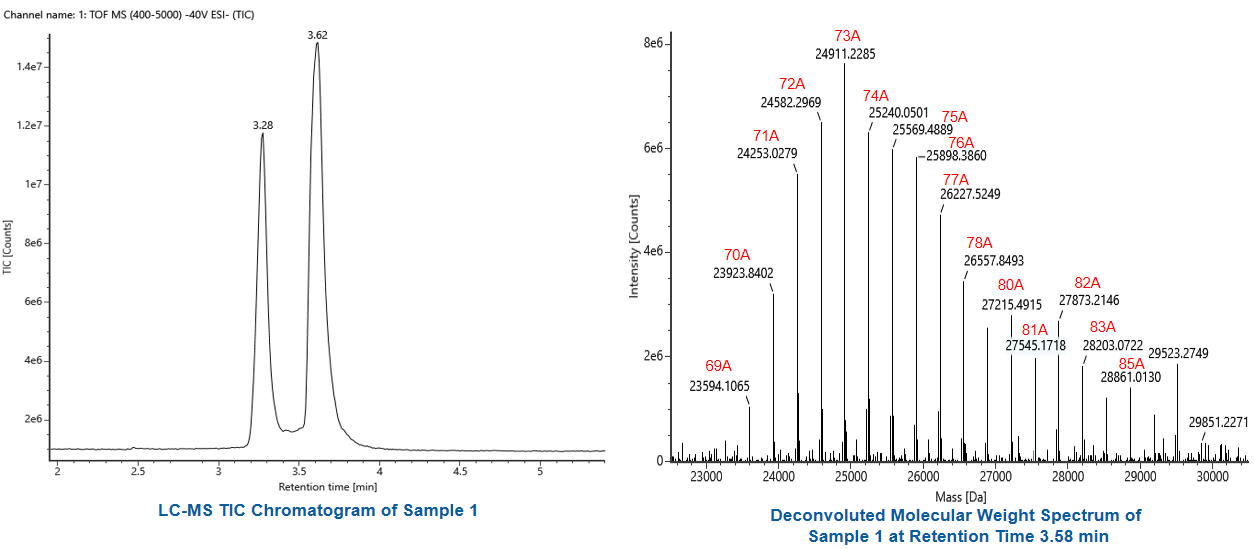 |
Total protein residue | Nanoorange | 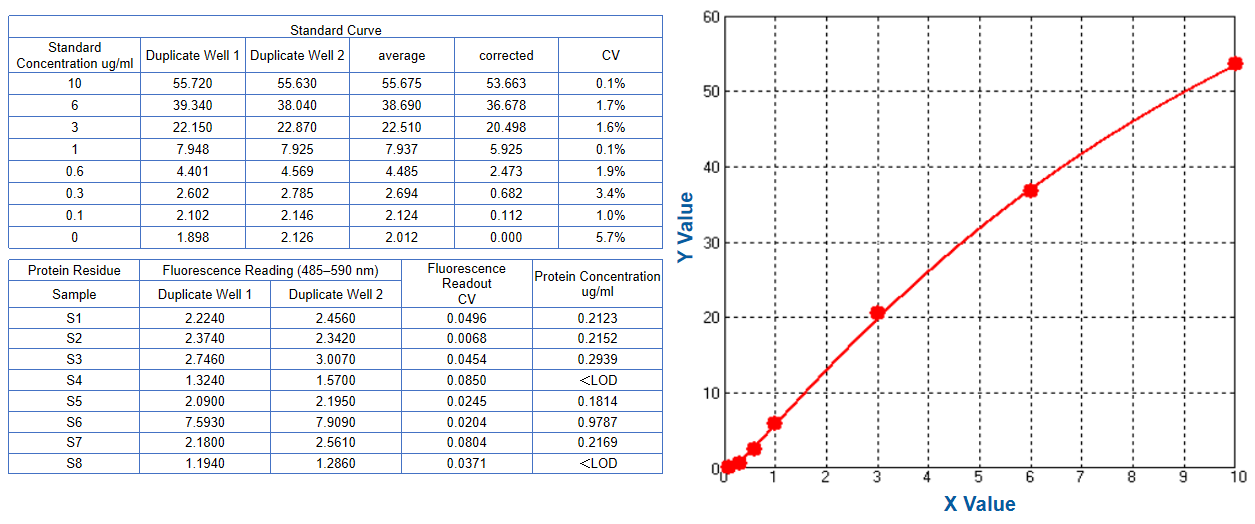 |
Product related impurities - NTP residue | SEC-HPLC | 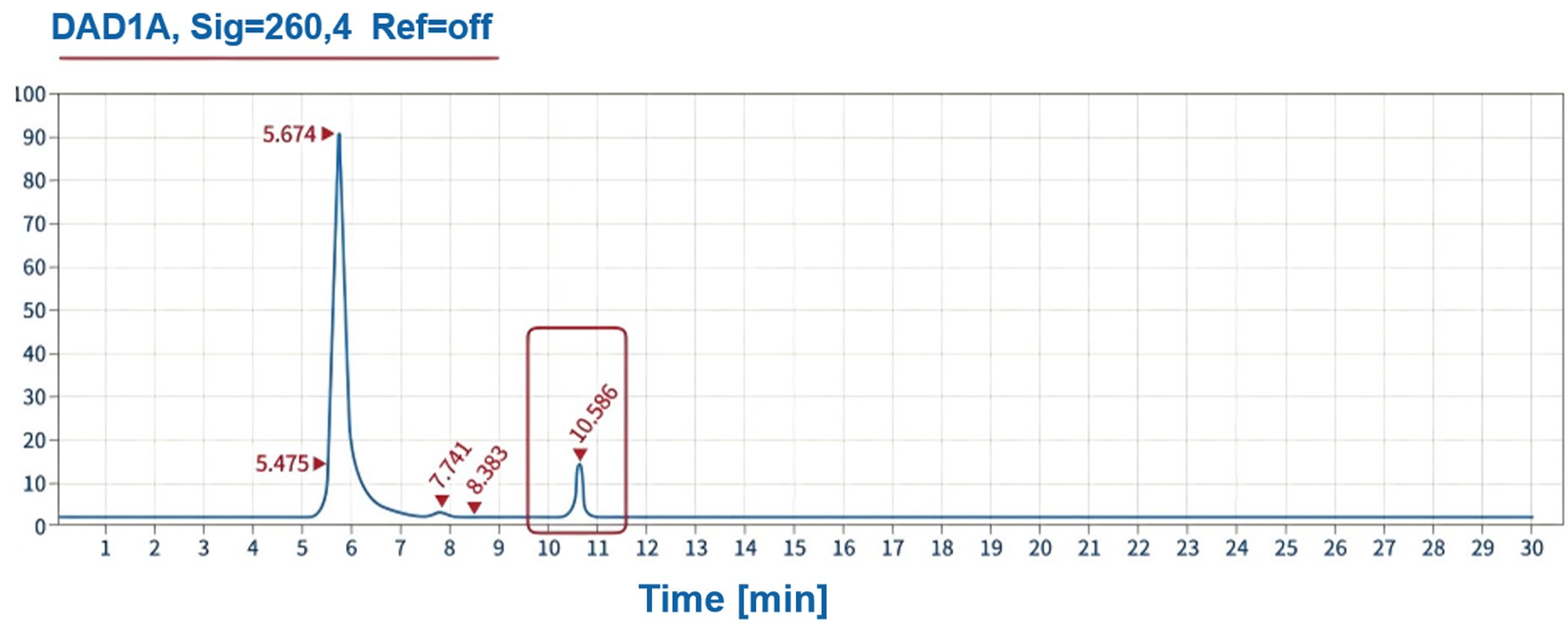 |
Product related impurities - polymer quantification | SEC-HPLC | 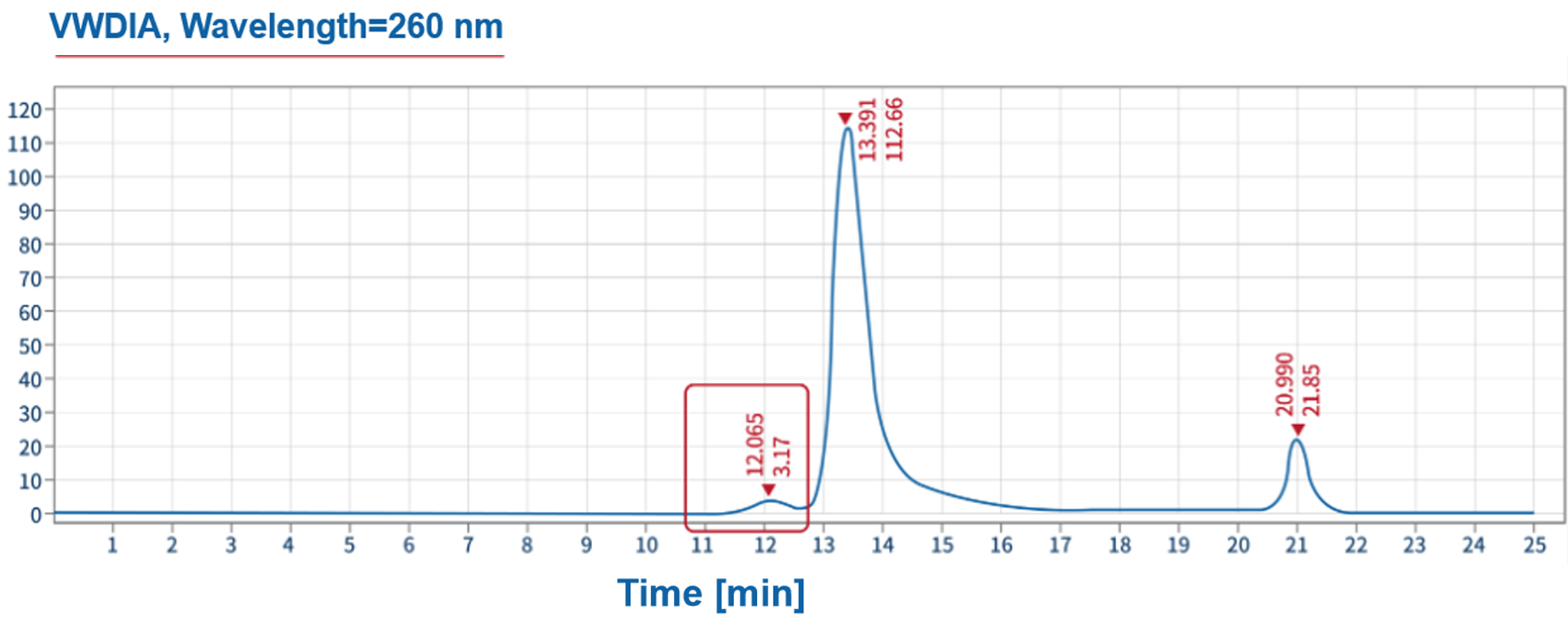 |
Product related impurities - residual T7 RNA polymerase content | ELISA | 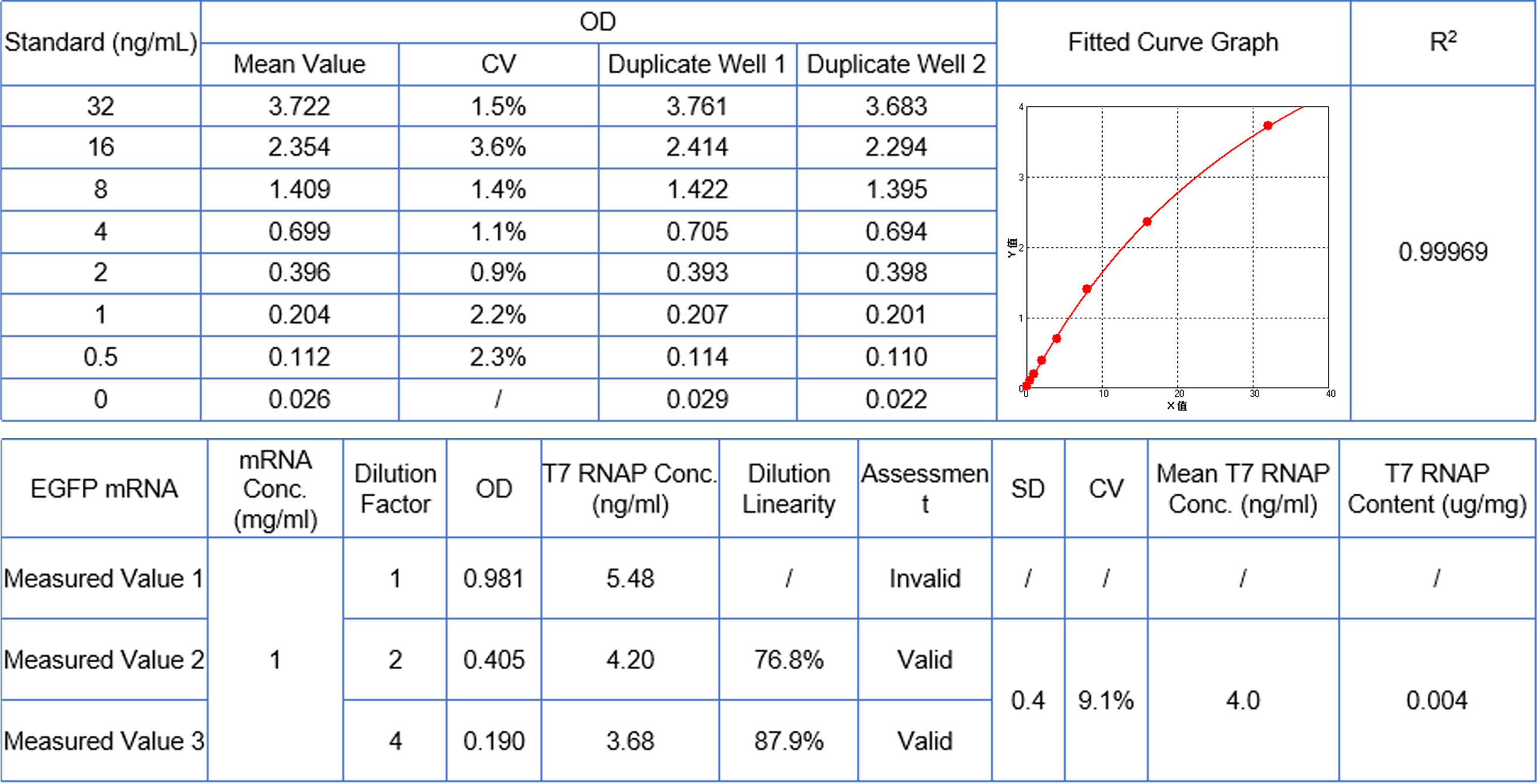 |
Endotoxin | Recombinant factor C/TAL |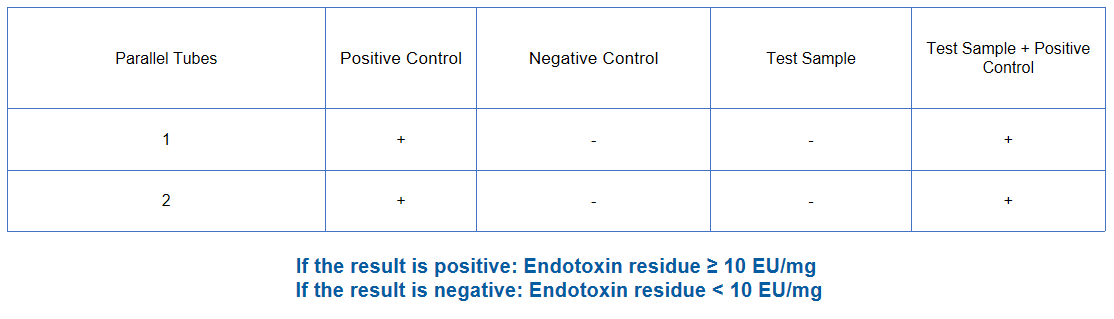  |
Verification level | Test method | Cycle | Results Display |
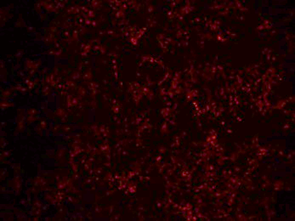
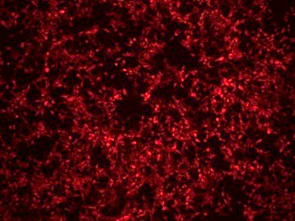
Verification of intracellular expression levels | Fluorescence microscopy detection | 3 days | 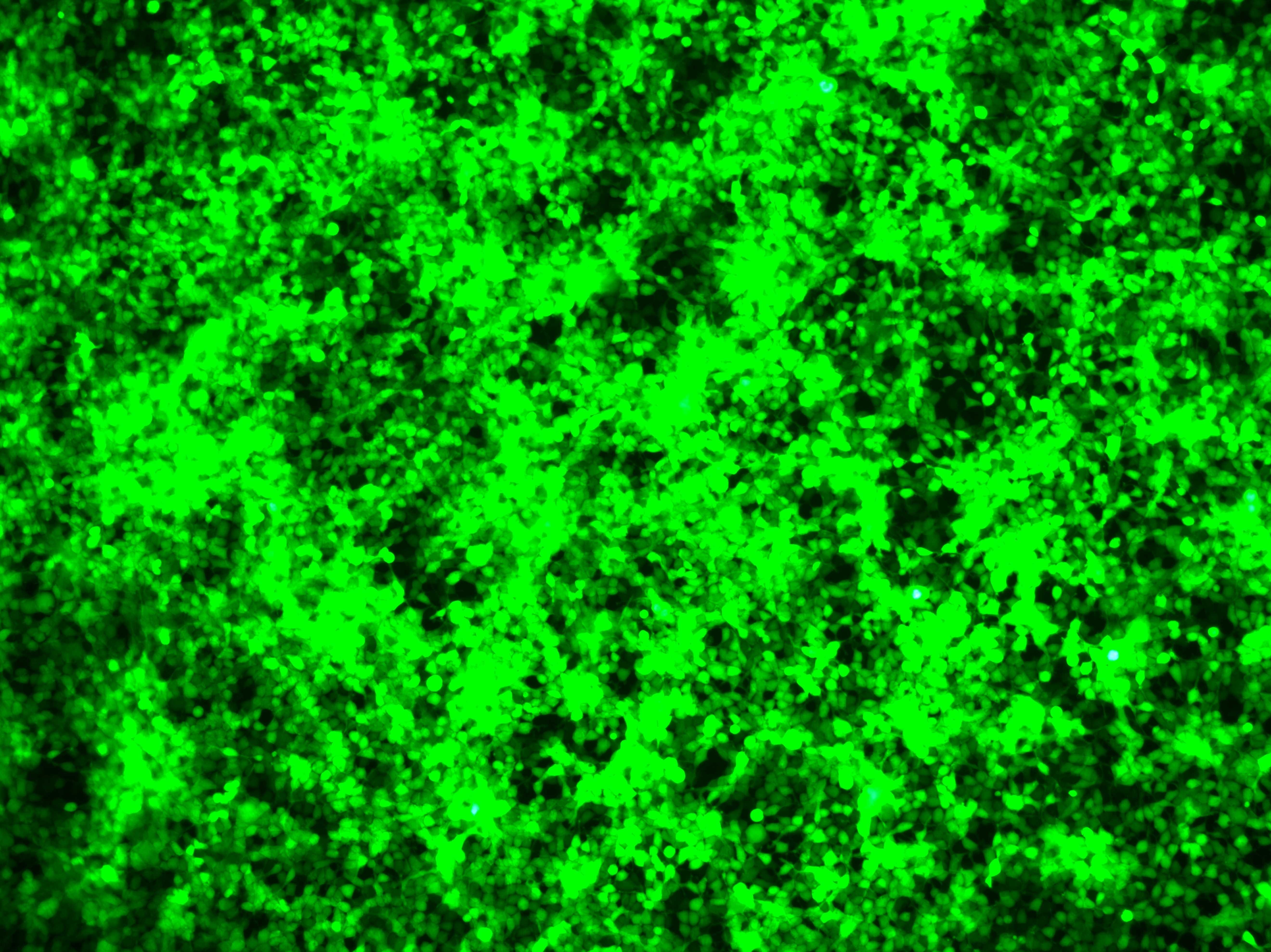 |
Flow cytometry analysis | 3 days | 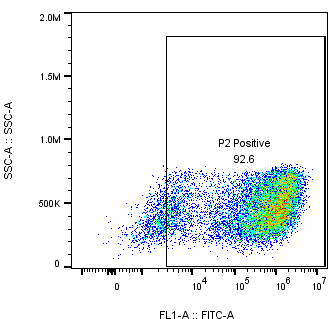 | |
Detection of firefly luciferase | 3 days | 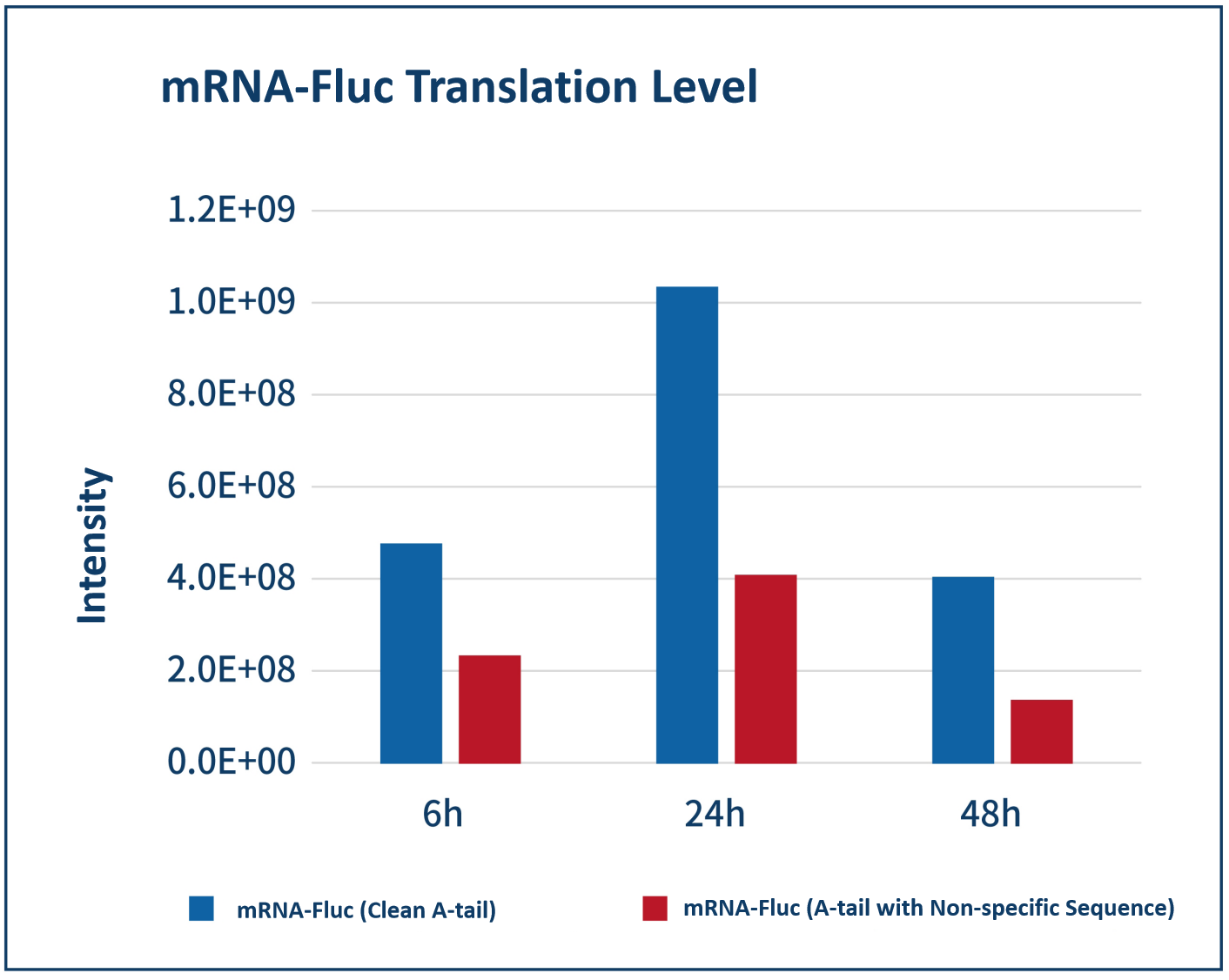 | |
Immunofluorescence | 3 days | 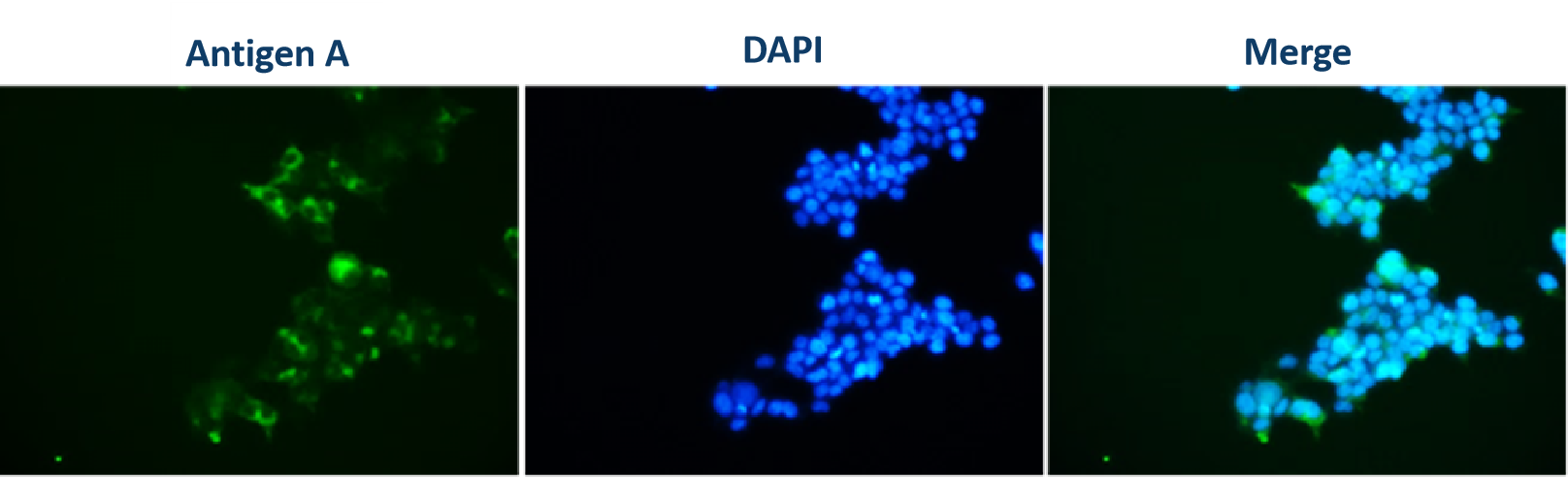 | |
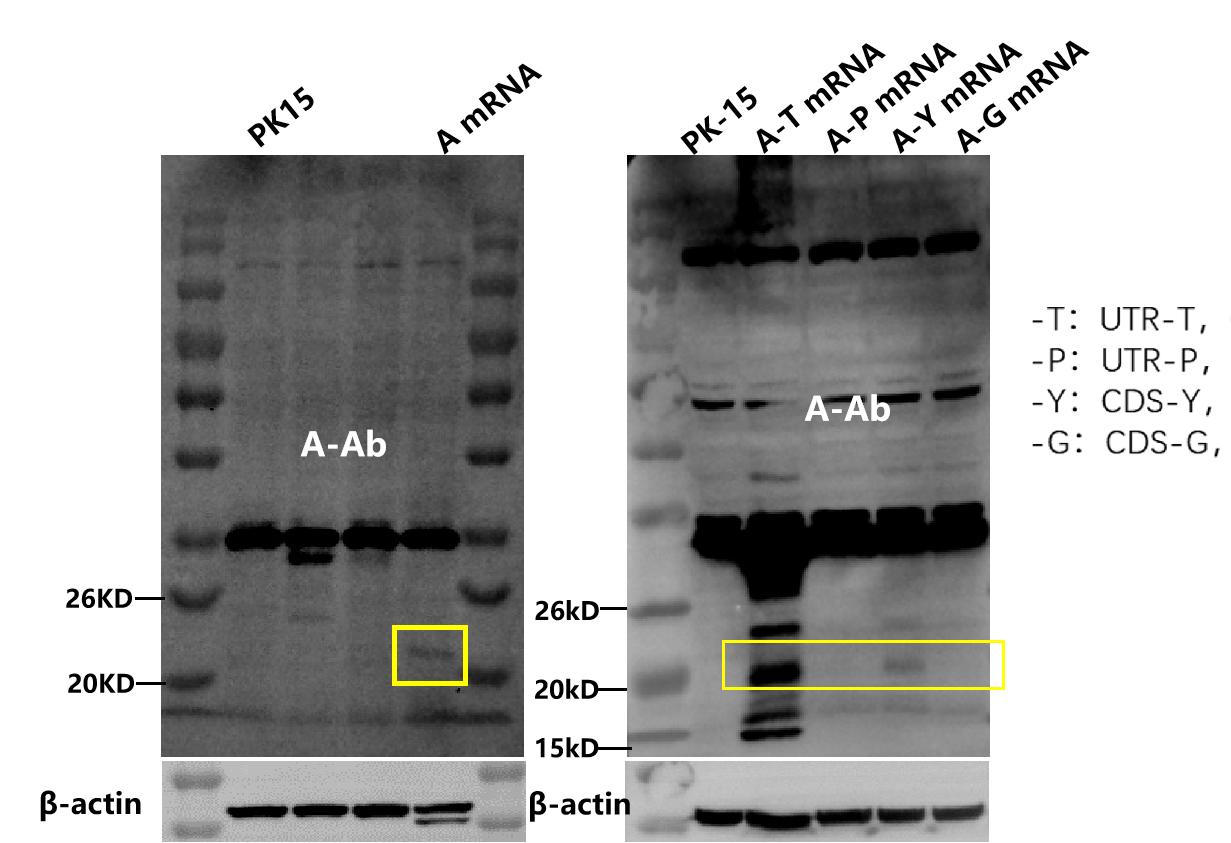
WB | 3 days | 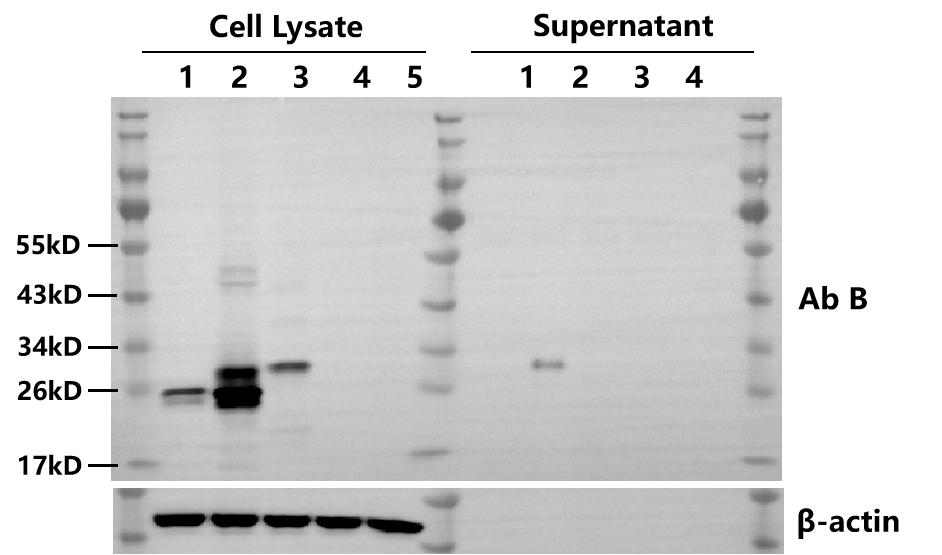 | |
ELISA | 3 days | 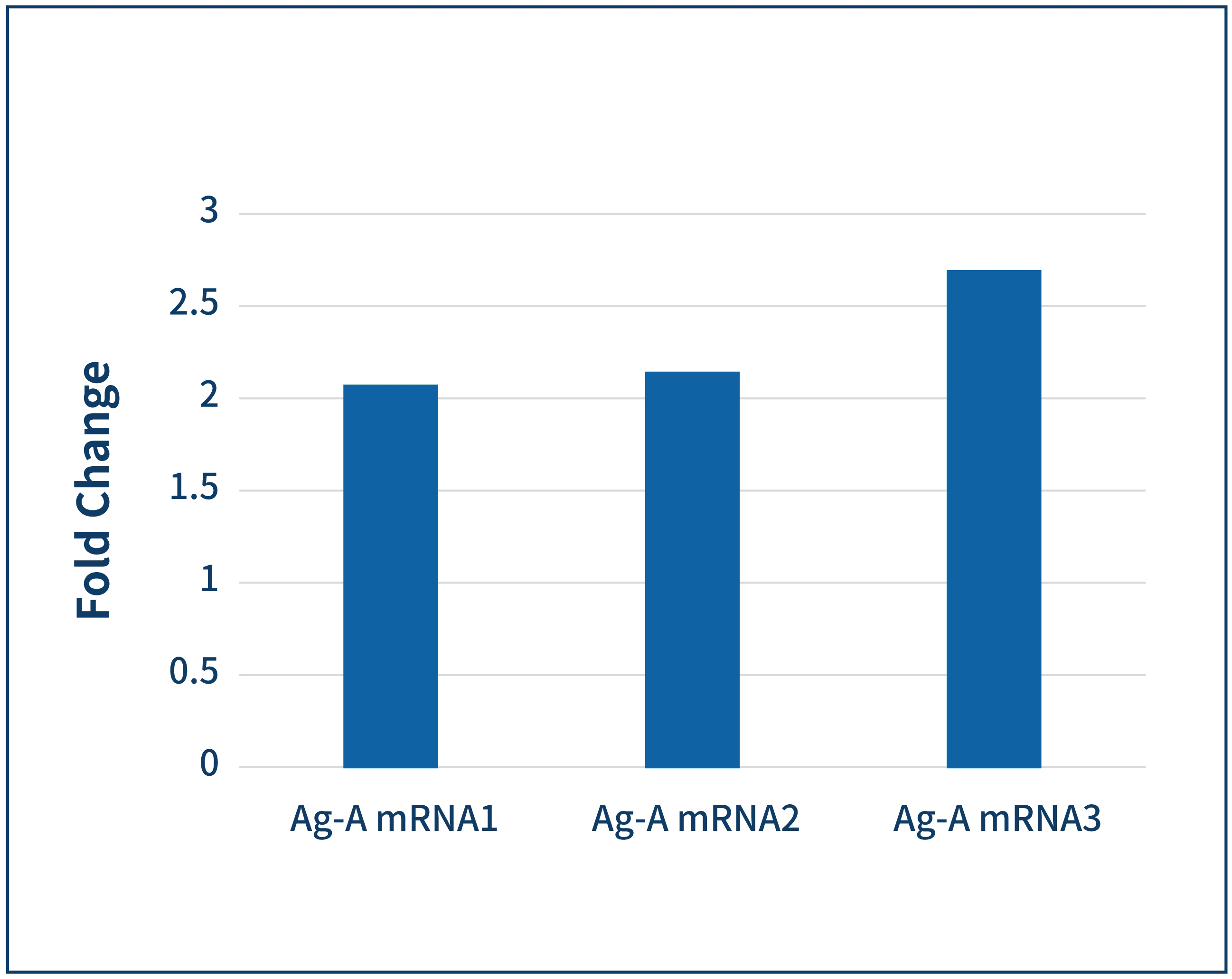 | |
In vivo expression level validation | In vivo imaging | 1-2 days | 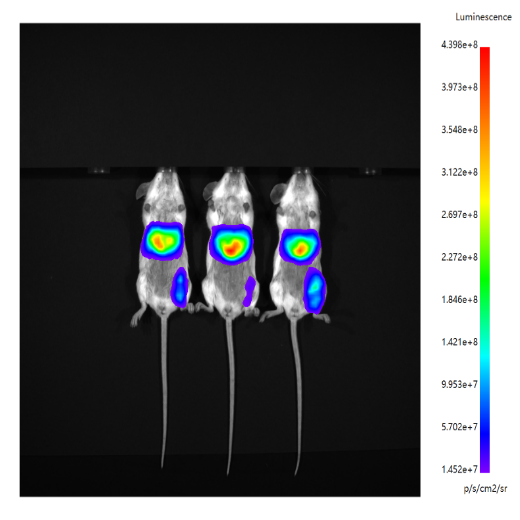 |
Serum protein expression detection | 2 days | 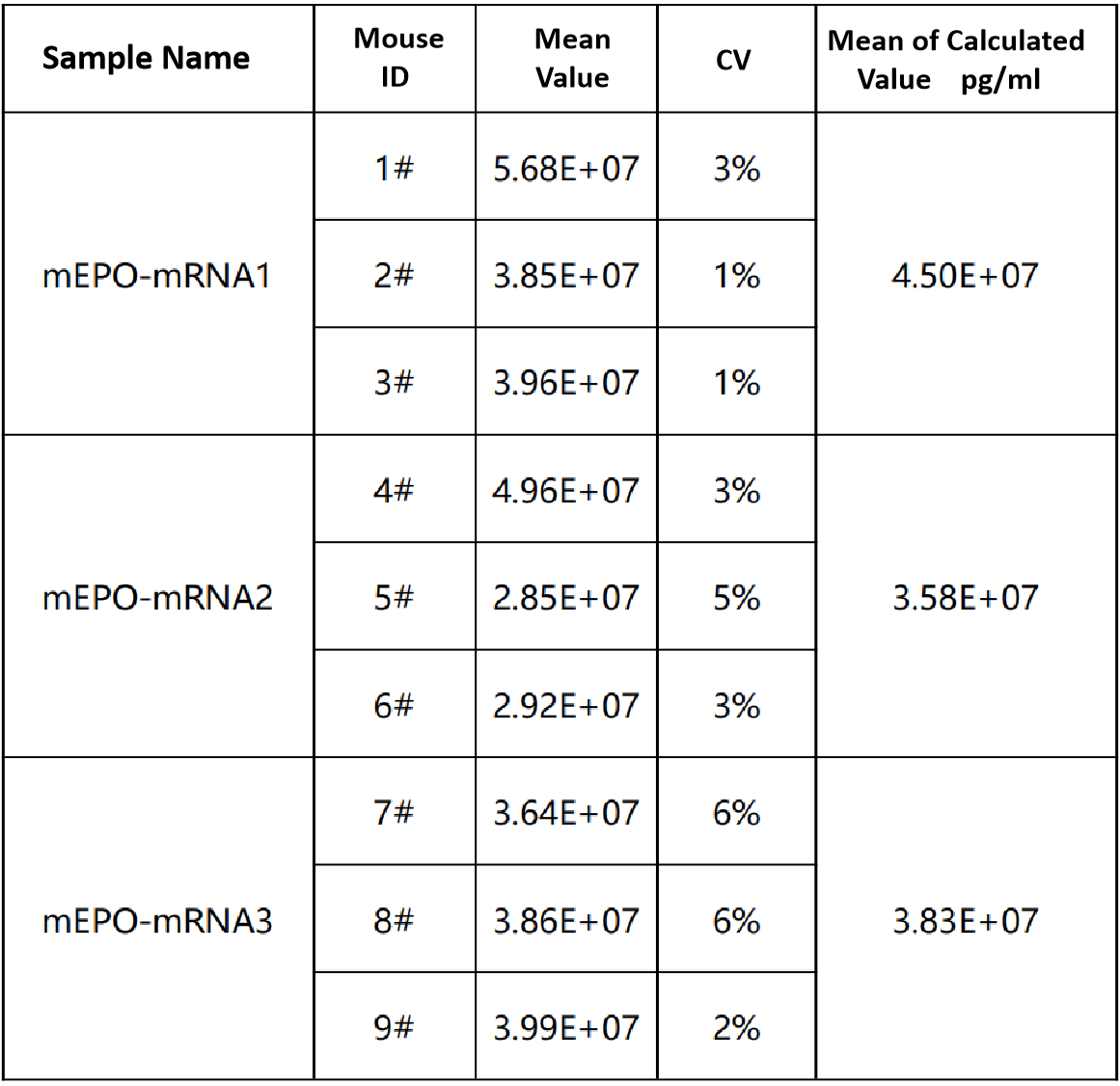 |

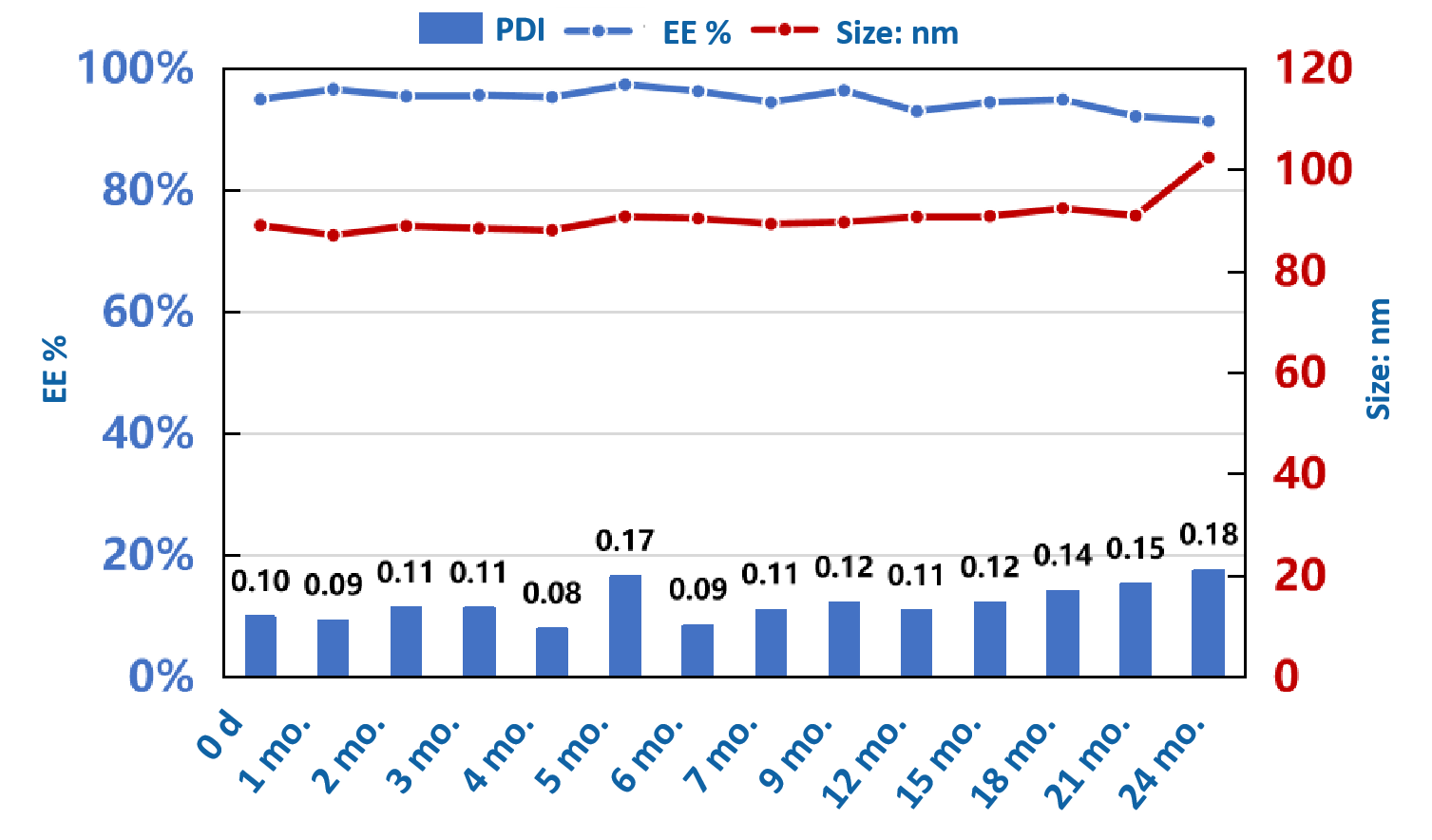
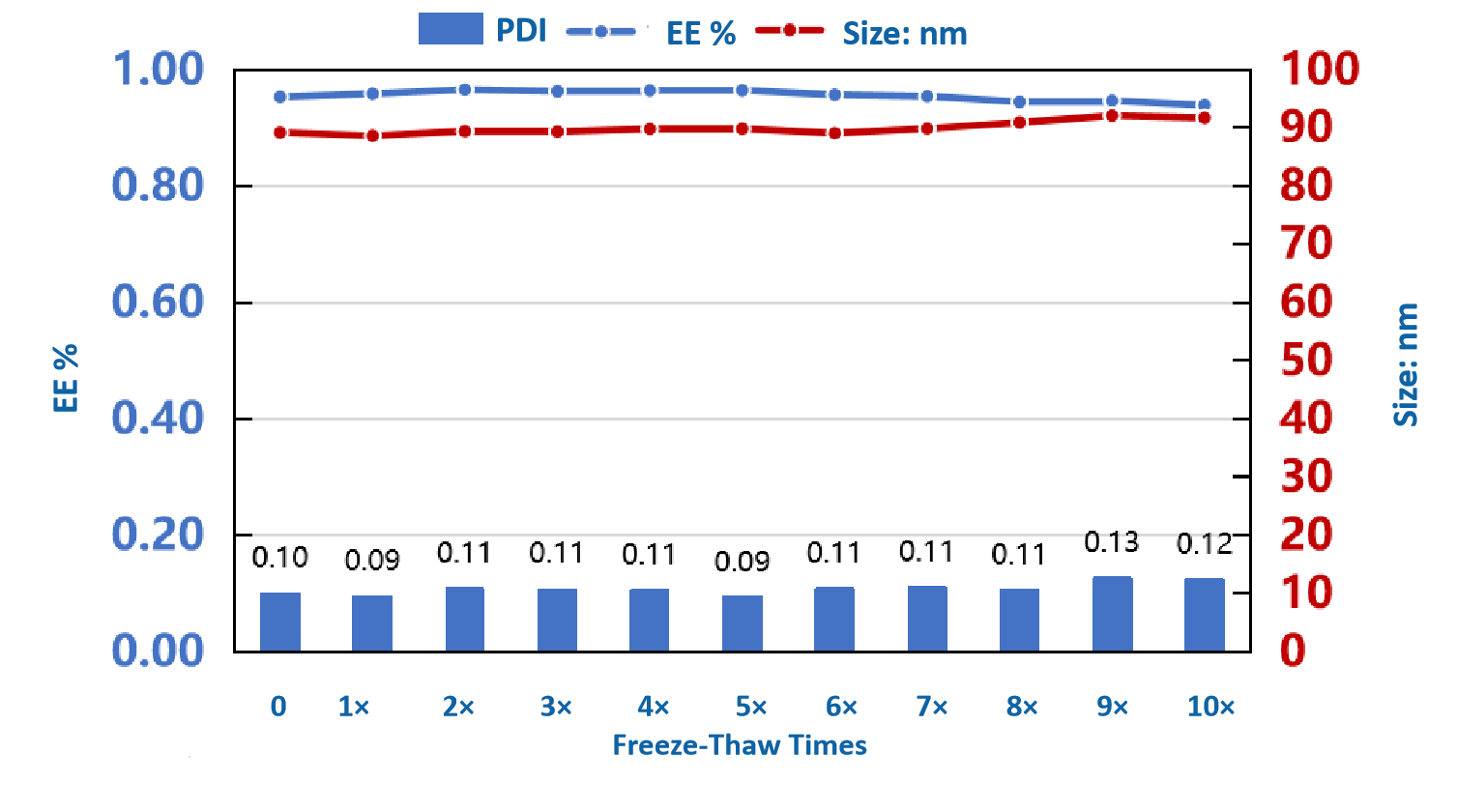
Product release testing
Quality inspection items | Quality inspection method | Results Display |
Entrapment efficiency | Ribogreen | 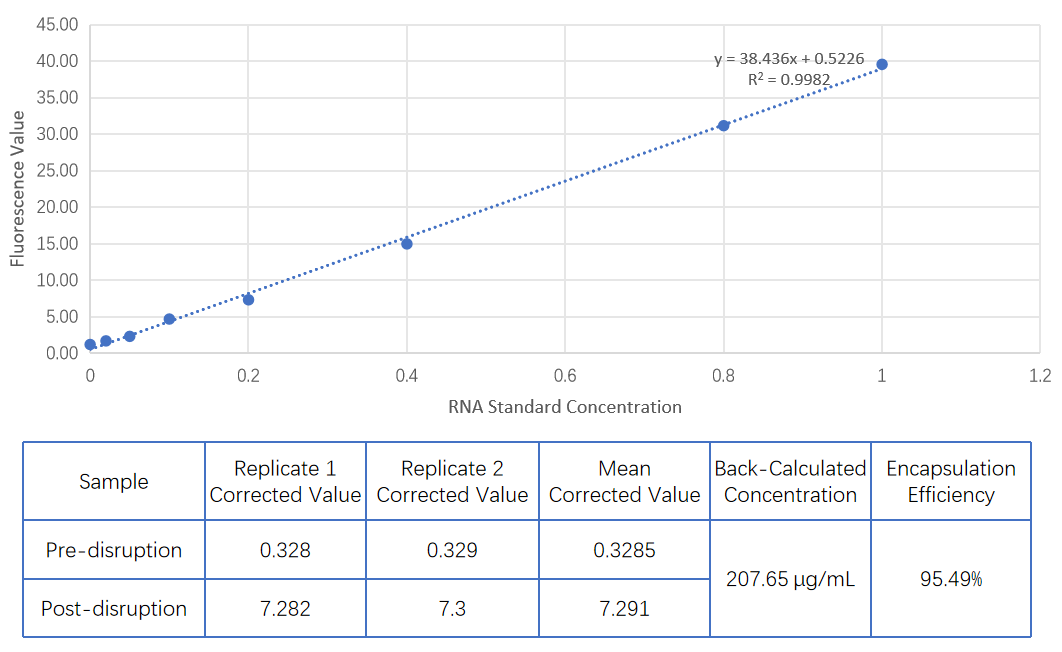 |
Preparation RNA concentration | ||
PDI/particle size | DLS | 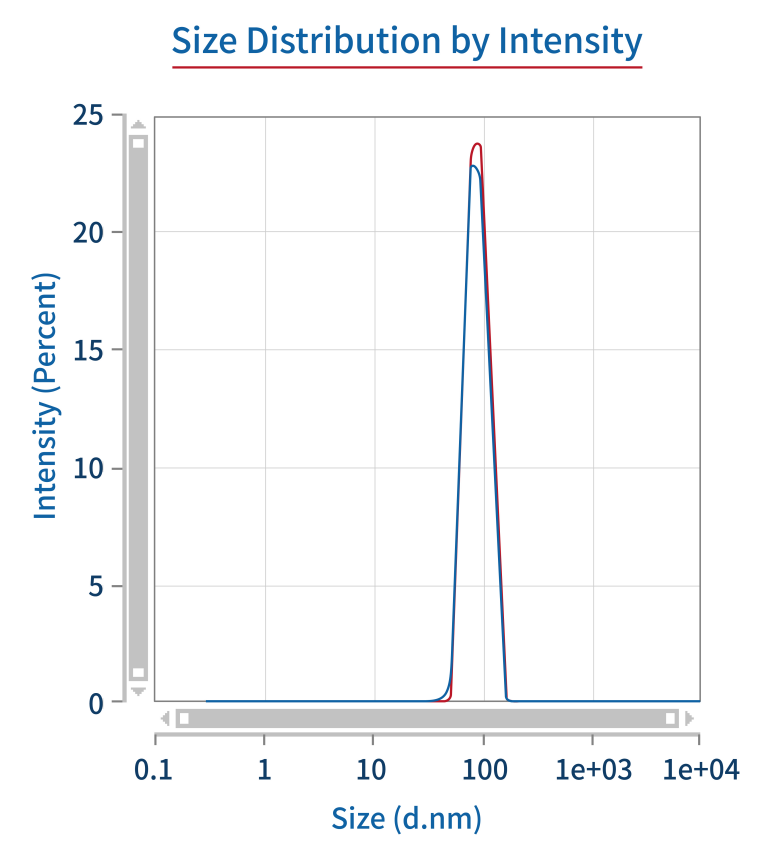 |
Zeta potentials | Doppler electrophoresis | 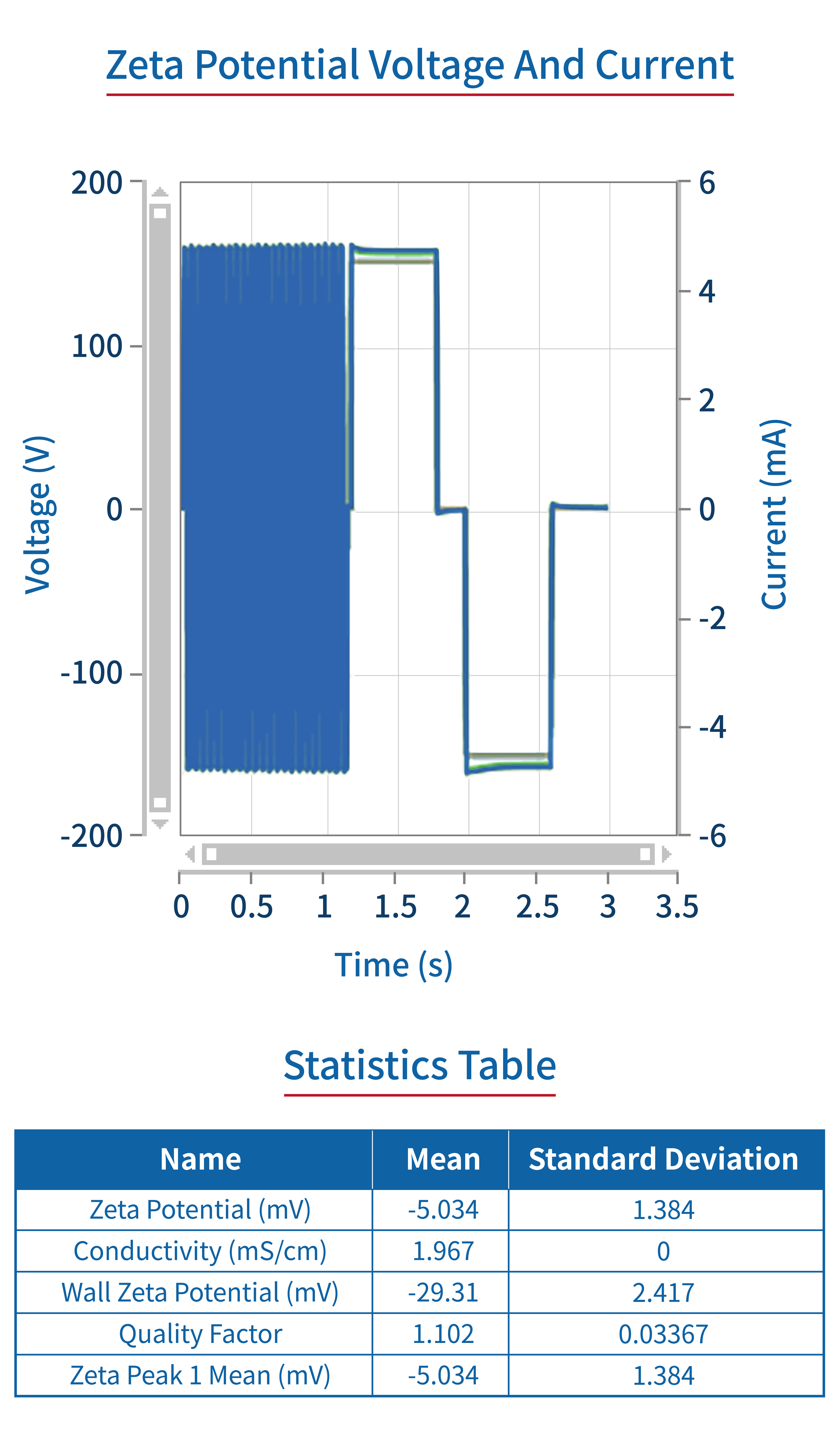 |
Molecular molar ratio of multivalent RNA molecules | ddPCR+qPCR | 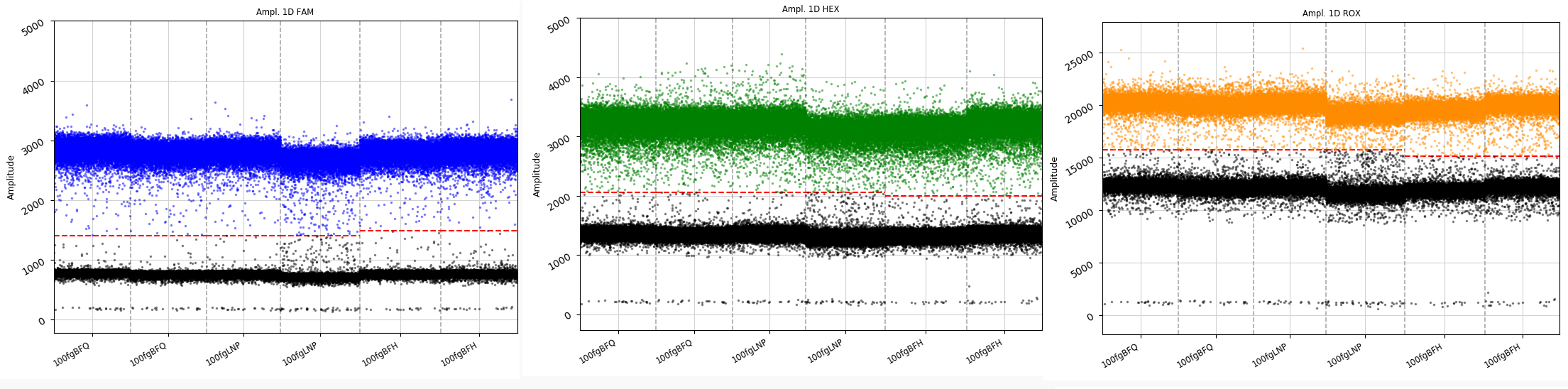 |
RNA sequence confirmation | Sanger sequencing/RT-PCR | 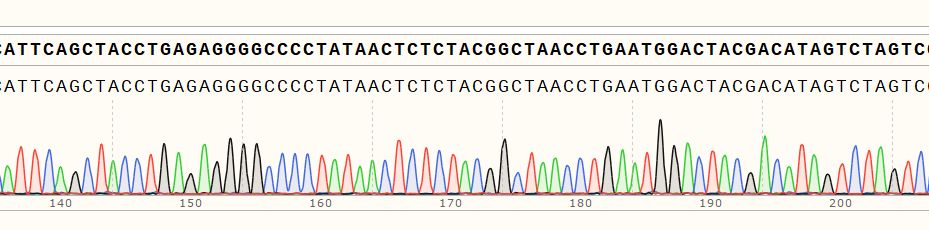 |
Determination of lipid content | ELSD-HPLC | 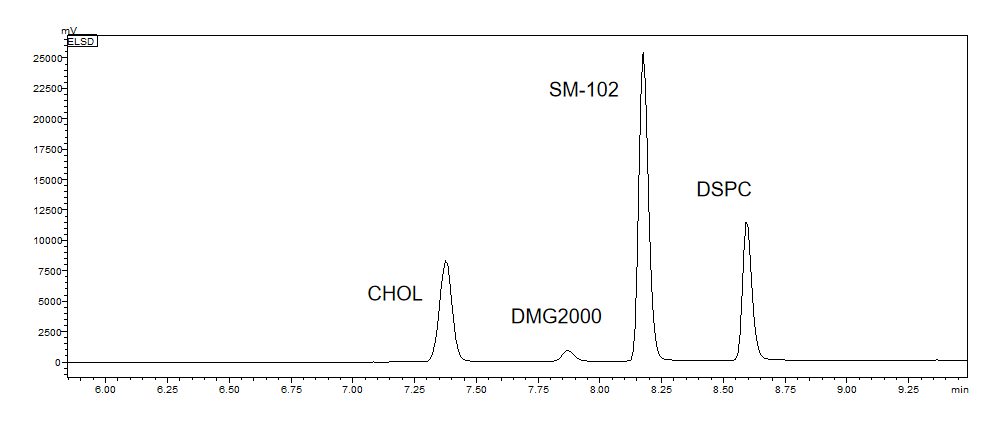 |
RNA integrity | CE/HPLC | 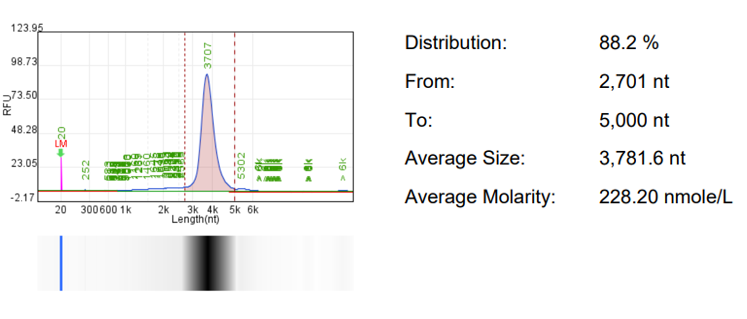 |
Endotoxin | Recombinant factor C/TAL | 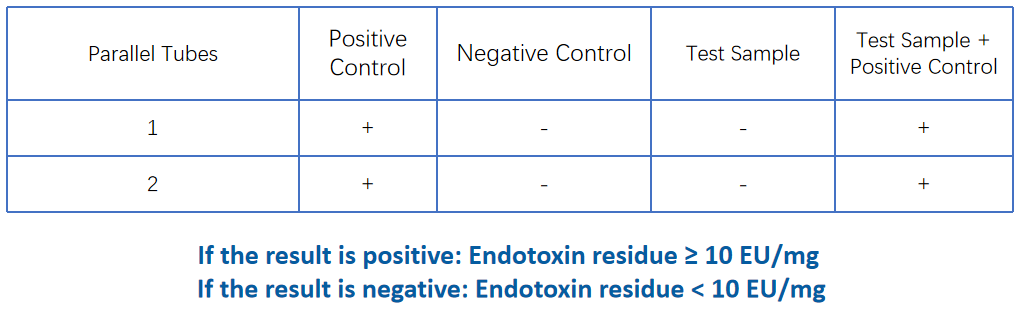 |
Protein expression validation | WB/FACS/IF/ELISA |


